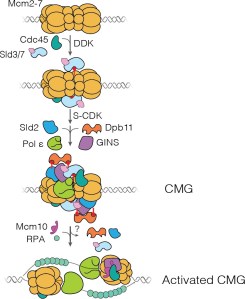
 Replication of chromosomal DNA in eukaryotes has two major stages. Starting in the G1 phase of the cell cycle, double hexamers consisting of two copies of the Mcm2-7 replicative helicase are assembled at replication origins. Later, in S phase, the two helicases are incorporated into two oppositely oriented CMG (Cdc45-Mcm2-7-GINS) complexes that each then form the core of a replisome. Control of this “activation” step, which is triggered by the protein kinases DDK and S-CDK, is essential to ensure that each part of the genome is replicated once and only once in each cell cycle.
Replication of chromosomal DNA in eukaryotes has two major stages. Starting in the G1 phase of the cell cycle, double hexamers consisting of two copies of the Mcm2-7 replicative helicase are assembled at replication origins. Later, in S phase, the two helicases are incorporated into two oppositely oriented CMG (Cdc45-Mcm2-7-GINS) complexes that each then form the core of a replisome. Control of this “activation” step, which is triggered by the protein kinases DDK and S-CDK, is essential to ensure that each part of the genome is replicated once and only once in each cell cycle.
In this paper, together with collaborators from Steve Bell’s and Jeff Gelles’ labs used multi-wavelength single-molecule fluorescence colocation (“CoSMoS”) methods to study in vitro the molecular mechanism of the activation process. The journal’s acceptance summary notes that “The manuscript provides new and convincing evidence that a heretofore unknown intermediate state [called “CtG”] for replication start contains multiple copies of the GINS and Cdc45 proteins prior to initiation at each origin with one double hexamer of the MCM2-7 complex. The number of GINS and Cdc45 is determined by DDK phosphorylation of the MCM’s and the probability to create an active helicase (CMG) is increased with multiple numbers of the bound ancillary factors…. The single molecule studies and biochemistry are beautifully executed providing the evidence for such intermediates…. The addition of in vivo studies demonstrates that modulating the multiplicity of DDK phosphorylation (and proposed, CtG formation) has an impact on origin usage in cells.”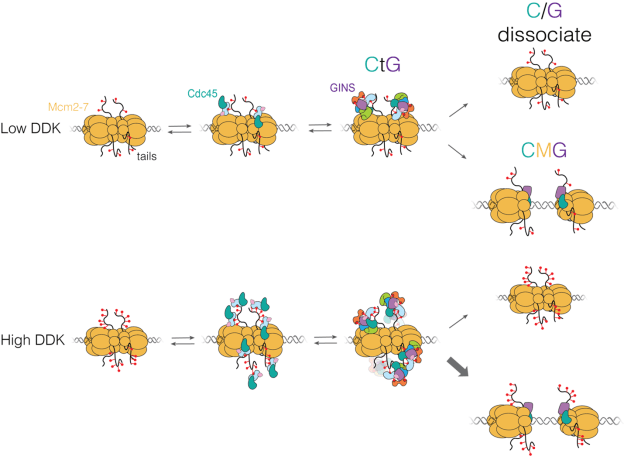
Kim L.D.J., et al., DDK regulates replication initiation by controlling the multiplicity of Cdc45-GINS binding to Mcm2-7.
eLife 10, e65471 (2021)

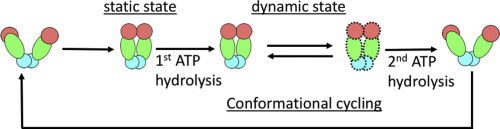
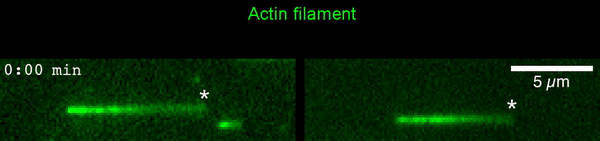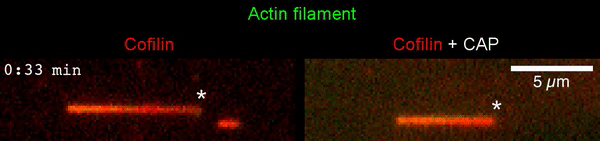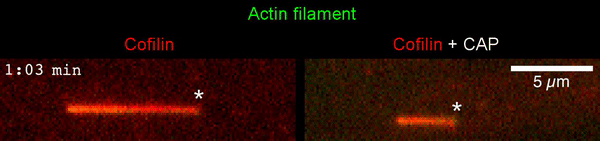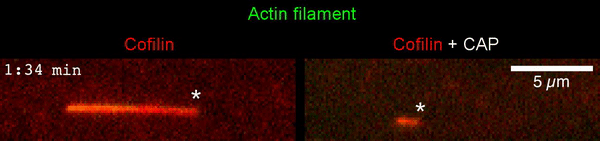 The new study shows that Cofilin and one other protein (Srv2/CAP) intimately collaborate at one end of the actin filament to accelerate subunit dissociation by over 300-fold! These are the fastest rates of actin depolymerization ever observed. Further, these results establish a new paradigm in which a protein that decorates filament sides (Cofilin) works in concert with a protein that binds to filament ends (Srv2/CAP) to produce an activity that is orders of magnitude stronger than the that of either protein alone.”
The new study shows that Cofilin and one other protein (Srv2/CAP) intimately collaborate at one end of the actin filament to accelerate subunit dissociation by over 300-fold! These are the fastest rates of actin depolymerization ever observed. Further, these results establish a new paradigm in which a protein that decorates filament sides (Cofilin) works in concert with a protein that binds to filament ends (Srv2/CAP) to produce an activity that is orders of magnitude stronger than the that of either protein alone.”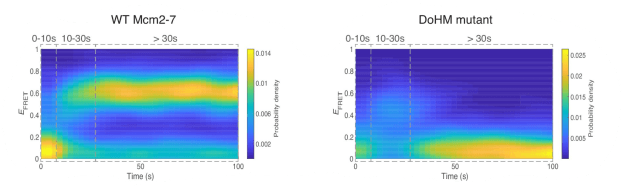
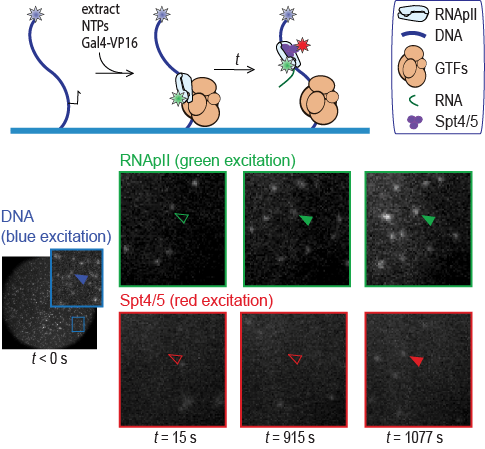
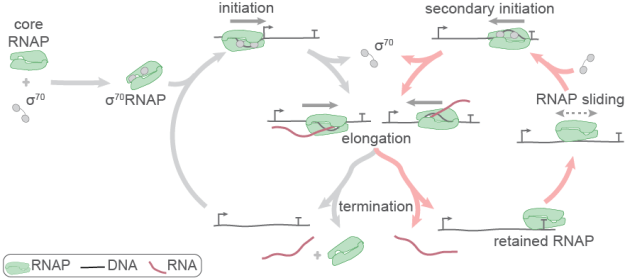
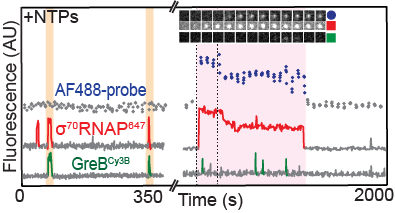 Sarah Stumper and her collaborators used fluorescence correlation spectroscopy and multi-wavelength single-molecule fluorescence colocalization microscopy to show that the SCFs likely repress transcription through an interesting “delayed inhibition” mechanism in which the proteins arrive at DNA already complexed to RNA polymerase and block at a later stage of transcription initiation. The work explains factors that control the relative contributions of the two proteins to regulation and suggests a mechanism by which repression is restricted to ribosomal RNA and other promoters that form short-duration complexes with RNA polymerase.
Sarah Stumper and her collaborators used fluorescence correlation spectroscopy and multi-wavelength single-molecule fluorescence colocalization microscopy to show that the SCFs likely repress transcription through an interesting “delayed inhibition” mechanism in which the proteins arrive at DNA already complexed to RNA polymerase and block at a later stage of transcription initiation. The work explains factors that control the relative contributions of the two proteins to regulation and suggests a mechanism by which repression is restricted to ribosomal RNA and other promoters that form short-duration complexes with RNA polymerase.